Abstract
Checkpoint kinase 1 inhibitors (CHK1i) have shown impressive single-agent efficacy in treatment of certain tumors, as monotherapy or potentiators of chemotherapy in clinical trials, but the sensitive tumor types and downstream effectors to dictate the therapeutic responses to CHK1i remains unclear. In this study we first analyzed GDSC (Genomics of Drug Sensitivity in Cancer) and DepMap database and disclosed that hematologic malignancies (HMs) were relatively sensitive to CHK1i or CHK1 knockdown. This notion was confirmed by examining PY34, a new and potent in-house selective CHK1i, which exhibited potent anti-HM effect in vitro and in vivo, as single agent. We demonstrated that the downregulation of c-Myc and its signaling pathway was the common transcriptomic profiling response of sensitive HM cell lines to PY34, whereas overexpressing c-Myc could partially rescue the anticancer effect of PY34. Strikingly, we revealed the significant correlations between downregulation of c-Myc and cell sensitivity to PY34 in 17 HM cell lines and 39 patient-derived cell (PDC) samples. Thus, our results demonstrate that HMs are more sensitive to CHK1i than solid tumors, and c-Myc downregulation could represent the CHK1i efficacy in HMs.
Similar content being viewed by others
Introduction
The important functional roles of CHK1 in the DNA damage response (DDR) and cell cycle checkpoints were clarified in the 1990s [1, 2]. CHK1 is overexpressed in a variety of solid tumors and hematologic malignancies (HMs) [3, 4]. CHK1 siRNA or CHK1 inhibitors (CHK1is) had little effect on cell viability in healthy peripheral blood mononuclear cells [5] but showed significant cellular cytotoxicity in neuroblastoma [6], chemical carcinogen-induced mouse skin tumorigenesis [7] and chronic lymphocytic leukemia cells [5], which implied that CHK1 is essential for tumor cell survival but redundant for normal cells [8, 9]. Furthermore, CHK1 activation may also contribute to therapeutic resistance in a wide variety of tumor treatments, including chemotherapy, radiotherapy, targeted therapy, and immunotherapy. For example, increased expression of CHK1 could enhance chemoresistance in acute myeloid leukemia (AML) [10], promote cisplatin resistance in ovarian cancer [11], predict early local recurrence and radioresistance in breast cancer [12], contribute to oxaliplatin resistance in colorectal cancer [13], and elicit PD-L1 immune checkpoint activation [14]. For the reasons mentioned above, CHK1 is a promising cancer therapeutic target.
Until now, in total, 11 CHK1is have entered clinical trials against solid tumors or HMs and have been used as mono- or combination therapy in phase I or phase II clinical trials [15]. However, it remains unknown which tumor types are sensitive to CHK1is and whether there exists a key downstream effector that could indicate the effectiveness of CHK1i, which is important for directing the use of CHK1i in clinical treatment.
Here, in silico data mining of the GDSC (Genomics of Drug Sensitivity in Cancer) [16] and DepMap [17] databases was conducted, which revealed that HMs were more sensitive to CHK1i or CHK1 knockdown than solid tumors. Furthermore, PY34, a new and potent in-house selective CHK1i [18], showed good anti-HM effects as a single agent in vitro and in vivo. Mechanistically, the expression of c-Myc and its downstream genes was significantly downregulated in sensitive cells after PY34 treatment via transcriptional repression. Overexpressing c-Myc partially reversed the inhibition of cell proliferation induced by PY34. In addition, c-Myc downregulation was significantly associated with the sensitivity of the HM cell line panel and AML PDC samples to PY34. Our findings suggested that HMs will most likely respond to CHK1i monotherapy and that c-Myc downregulation is a candidate indicator of CHK1i efficacy in HMs, which merits further investigation in follow-up preclinical studies and human clinical trials.
Materials and methods
Data mining and analysis of the GDSC database
The GDSC database currently contains data for 187 drugs and more than 160,000 IC50 values (half maximal inhibitory concentration). Two datasets are available: GDSC1 updates previous screening results (Resazurin or Syto60 Assay), and GDSC2 uses the most recently developed screening technology (CellTitreGlo Assay). The public online platform (https://www.cancerrxgene.org/) was developed by GDSC, in which all the data are downloadable and plots are reproducible. The data encompassing the sensitivities of a variety of cancer cell lines to CHK1is were downloaded, and scatter plots and either unpaired t-tests or Mann–Whitney tests were generated and computed by GraphPad Prism 7.
Data mining and analysis of the DepMap database
Project Achilles is a systematic effort aimed at identifying and cataloging gene essentiality across hundreds of genomically characterized cancer cell lines whose data are hosted on the Cancer Dependency Map Portal (DepMap, https://depmap.org/portal/). A lower score means that a gene is more likely to be dependent in a given cell line. The Genetic Dependency dataset (CRISPR (Avana) Public 19Q4 version) was downloaded, and then the corresponding gene-dependence scores for different cell lines were obtained by looking up gene names (such as CHEK1 and MYC). Scatter plots, unpaired t-tests or Mann-Whitney tests, and correlation analyses were generated by GraphPad Prism 7.
Cell culture, transduction, and puromycin selection
Cells were maintained in the appropriate culture medium suggested by suppliers (Supplementary Table S1). All cells were grown in the presence of 100 U/mL penicillin and 100 μg/mL streptomycin at 37 °C and 5% CO2. All cell lines were used within 15 passages and less than 6 months.
For CHK1 knockdown, GV307, an miR30-based shRNA vector [19] was used. Vectors containing control shRNA (5′-TCTCGCTTGGGCGAGAGTAAG-3′) or CHK1 shRNA (5′-GCAACAGTATTTCGGTATAAT-3′) and lentivirus were purchased from GeneChem Group (Shanghai, China). MV411, Z138, and K562 cells were infected with lentivirus in the presence of 6 mg/mL polybrene and selected with puromycin for 2 weeks. Doxycycline (1 μg/mL) was added to induce shRNA expression to knockdown CHK1. For c-Myc rescue experiments, the retroviral constructs MSCV-mCherry-c-Myc and MSCV-mCherry-empty vectors were gifted from the School of Medicine, Shanghai Jiao Tong University. To generate cells with stable expression, the plasmids were transfected into Lenti-X™ 293T (Clontech, Mountain View, CA, USA) packaging cells with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). After 48 h, medium containing virus was collected, filtered, and used to infect host cells in the presence of 6 mg/mL polybrene. Stable transfectants with mCherry fluorescence were sorted by flow cytometry.
In vitro antiproliferative activities
Antiproliferative activities were assessed by the MTS assay. Briefly, cells were cultured in triplicate for 3 days with the compounds tested. Then, CellTiter 96® Aqueous Non-Radioactive Cell Proliferation Assay Reagent (Promega Corporation, Madison, WI, USA) was added 3 h before the end of culture. The OD490 was measured with a 96-well plate reader. The IC50 value of the compound was calculated by GraphPad Prism 7.
Cell cycle analysis
Cells were harvested and fixed in ice-cold 70% ethanol overnight and then stained with propidium iodide (P4170, Sigma-Aldrich LLC, Darmstadt, Germany). Data were collected on a BD Accuri™ C6 flow cytometer (Becton, Dickinson and Company (BD), Franklin Lakes, NJ, USA) and analyzed with FlowJo software.
Transcriptomics profiling analysis
Sequencing was performed by Novogene Bioinformatics Technology Co., Ltd (Beijing, China) and the HISAT2, Samtools, Htseq-count, and DEseq2 packages in R language software were used to analyze the differential genes. Online tools or R language packages, such as Jvenn, Metascape, pheatmap, and GSEA, were used for subsequent analysis. Briefly, RNA-seq data were analyzed using the following strategy. Jvenn was used to overlap the differentially expressed genes. Based on the hallmark database, Metascape was used to enrich the differentially expressed genes. The pheatmap package in R language was used to analyze the expression differences. Gene set enrichment analysis (GSEA) was used to detect changes in the transcription factor pathways based on the transcription factor target gene sets.
Western blot
Cells were lysed in 100 μL of Laemmli sample buffer (#1610747, Bio-Rad Laboratories, Hercules, CA, USA) and boiled for 15 min. Proteins were then resolved via SDS-PAGE and transferred to nitrocellulose membranes (#10600002, GE Healthcare, Fairfield, CT, USA). After blocking with 5% nonfat milk or 5% bovine serum albumin, membranes were blotted with the antibodies listed in Supplementary Table S2. Detection was achieved with an Odyssey® Imaging System (LI-COR Biosciences, Lincoln, NE, USA), and the band intensities were quantified by ImageJ software.
RNA extraction, reverse transcription, and quantitative PCR (qPCR)
RNA was harvested with an RNAiso Plus kit (#9108, Takara Biomedical Technology, Beijing, China), and first-strand cDNA synthesis was performed with the PrimeScript™ RT Master Mix kit (Q131, Vazyme Biotech, Nanjing, China) according to the manufacturers’ specifications. Samples were analyzed on a Stratagene MX3005P™ PCR instrument (Agilent Technologies, Santa Clara, CA, USA) using the ΔΔCT method. The primer sequences used are shown in Supplementary Table S3.
Xenograft model evaluation
Female NOD-SCID mice or NU/NU mice (7–8 weeks old) were purchased from Vital River Laboratory Animal Technology Co., Ltd (Beijing, China). Animal use procedures were approved based on the guidelines established by the Committee for Laboratory Animal Research at the Shanghai Research Center for Model Organisms (IACUC approval numbers SD18058 and NN1804). Cells were resuspended in a serum-free growth medium and mixed 1:1 with Matrigel® Matrix (#354248, Corning, NY, USA). Mice were subcutaneously injected with 5 × 106 Z138 or MV411 cells in the right flank. After the median tumor volume reached 100–300 mm3, PY34 acetate or vehicle (5% mannitol) was administered intravenously (i.v.), and AraC was administered subcutaneously (sc) 5 times per week for 3 weeks. Tumor growth was monitored twice every week with digital caliper measurements, and body weight was measured. All animals were killed on day 22, and the tumors were harvested.
Patient samples and cell isolation
Bone marrow samples from patients with newly diagnosed and untreated AML, acute lymphoblastic leukemia (ALL), chronic myeloid leukemia or chronic lymphoblastic leukemia were analyzed. All patients were diagnosed with leukemia according to the 2016 revision of the World Health Organization classification of myeloid neoplasms and acute leukemia and were referred to our clinic at Changhai Hospital of Shanghai for morphology, cytochemistry, immunophenotyping, cytogenetics (karyotype and fluorescence in situ hybridization), and molecular analysis. The study was approved by the institutional review boards of Changhai Hospital affiliated with the Naval Medical University. Written informed consent from patients was obtained according to the Declaration of Helsinki. Mononuclear cells were isolated by density gradient centrifugation using Lymphoprep™ (#07851, STEMCELL Technologies, Oslo, Norway) and subjected to red blood cell lysis using lysis buffer (B541001, Sangon Biotech, Shanghai, China).
Statistical analysis
The difference between experimental groups in the in vitro and in vivo studies was compared using unpaired two-tailed Student’s t test analysis. P < 0.05 was considered statistically significant.
Results
HMs are relatively sensitive to CHK1i or CHK1 knockdown
To identify cancer types sensitive to CHK1i, the cell sensitivity data of two CHK1is, AZD7762 (datasets GDSC1 and GDSC2) and SCH900776 (dataset GDSC2), from the GDSC database were analyzed. For AZD7762, GDSC1 contains data from 158 HM cell lines and 787 solid tumor cell lines, and GDSC2 contains data from 131 and 633 cell lines. For SCH900776, GDSC2 contains data from 132 and 617 cell lines. Data from the three datasets showed that HMs, including leukemia, lymphoma, and myeloma, were more sensitive to AZD7762 or SCH900776 than solid tumors (Fig. 1a, b and Supplementary Fig. S1a, b). Furthermore, cell sensitivity to our in-house CHK1i, PY34, was tested at three doses on a panel of 60 cancer cell lines consisting of 27 HM and 33 solid tumor cells and then probed in a panel of 21 HM cell lines and 7 solid tumor cell lines for IC50 confirmation. HM cell lines were more sensitive to PY34 than solid tumors (Supplementary Fig. S1c, Fig. 1c). The antiproliferation IC50 of PY34 for most HM cell lines was below 1 μM, except for K562 and RPMI8226 cells. The top five most sensitive cell lines covered three types of HMs: lymphoma (Z138 and Ramos), leukemia (MV411 and CCRF-CEM), and multiple myeloma (MM1S). Clustering of the activities of PY34 and other positive CHK1is, including AZD7762, LY2606368, LY2603618, GDC-0575, SCH900776, and CCT245737, against six HM cell lines resulted in a stratification into two groups. AZD7762 and LY2606368, CHK1/CHK2 dual inhibitors and others, were highly selective CHK1is (Supplementary Fig. S1d).
The difference between HMs and solid tumors with regard to AZD7762 sensitivity based on data from 764 tumor cell lines from the GDSC2 dataset (a) and to SCH900776 based on data from 749 tumor cell lines from the GDSC2 dataset (b). c In vitro antiproliferative activities of PY34 on 29 cancer cell lines and the difference in these activities between HMs and solid tumors. d The CHK1 dependency score after CHK1 knockout from CRISPR (Avana) Public 19Q3 in the DepMap database. e The efficiency of knocking down CHK1 by qRT-PCR in Z138, MV411 and K562 cells with induction of shRNA by 1 μg/mL doxycycline (Doxy) for 4 days. The effect of CHK1 knockdown on the proliferation of Z138 (f), MV411 (g), and K562 (h) cells and the colony-forming ability of MV411 and K562 cells (i). *P < 0.05; **P < 0.01, versus vehicle group, unpaired two-tailed Student’s t test.
Furthermore, the proliferation of 58 HM cell lines was more dependent on CHK1 than that of 505 solid tumor cell lines, as indicated by the CHK1 dependence score after CHK1 knockout by CRISPR (Avana) Public 19Q3 in the DepMap database (Fig. 1d). Consistently, significant cell growth inhibition was observed in the CHK1i-sensitive MV411 and Z138 cells but not in the CHK1i-resistant K562 cells after CHK1 shRNA lentivirus treatment (Fig. 1e–h), and CHK1 shRNA reduced the colony formation capacity of MV411 cells but exerted no significant effect on K562 cells (Fig. 1i).
Targeting CHK1 is effective on HMs in vitro and in vivo
PY34, a new and potent in-house selective CHK1i [18], was used to evaluate the anti-HM effect in vitro and in vivo. PY34-induced divergent alterations in the cell cycle in the top three sensitive cells—G1 arrest in MV411 (Fig. 2a) and S phase arrest in Z138 and CCRF-CEM cells (Fig. 2b, c)—but exerted no effect on the cell cycle of insensitive cell line RPMI8226 (Supplementary Fig. S2a). A similar divergence in cell cycle arrest in sensitive cells was observed for another CHK1i, SCH900776 (Supplementary Fig. S2b, c). However, PY34 induced apoptosis in all three sensitive cancer cell lines in a dose-dependent manner (Fig. 2d–f). Accordingly, the anticancer effect of PY34 in mice subcutaneously inoculated with Z138, MV411, or CCRF-CEM cells was evaluated. PY34 alone significantly inhibited tumor growth and tumor weight in a dose-dependent manner without a significant loss in total body weight (Fig. 2g–o).
Cell cycle arrest of sensitive HM cell lines treated with PY34: MV411 (a), Z138 (b), and CCRF-CEM (c) cells were treated with PY34 for 24 h and analyzed by FACS. Apoptosis of PY34-sensitive HM cell lines: MV411 (d), Z138 (e), and CCRF-CEM (f) were treated with PY34 for 24 h and analyzed by FACS. g–o PY34 alone effectively inhibited the growth of MV411-, Z138-, and CCRF-CEM-xenografted tumors without affecting mouse weight: CB-17 SCID or NU/NU nude mice bearing MV411, Z138, and CCRF-CEM xenografts received 5% mannitol (vehicle control), PY34 or AraC once daily 5 days a week for 3 weeks (n = 8 for the treated group; n = 16 for the vehicle group). Tumor volumes (g, j, m) and total body weights (i, l, o) were measured twice a week, and the tumors were harvested and weighed on day 22 (h, k, n).
The expression of c-Myc and its related gene signature is downregulated in sensitive HM cells after PY34 treatment via transcriptional depression. Although MV411 and Z138 cells were arrested at different phases of the cell cycle after PY34 or SCH90076 treatment (Fig. 2a, b and Supplementary Fig. S2a, b), a common transcriptomic profiling response in Z138 and MV411 cells treated with PY34 was identified. Among the top 1000 genes modulated in the two cell lines, 208 were altered in both (Fig. 3a), and the c-Myc signaling pathway was the most enriched (Fig. 3b). GSEA independently confirmed changes in c-Myc-related gene signatures and gene list of Z138 and MV411 cells, where c-Myc expression was significantly downregulated (Fig. 3c, d). c-Myc has been proven to be a key downstream effector that determines the therapeutic response of some kinase-targeted cancer therapies [20, 21]. The downregulation of c-Myc expression by CHK1is was further confirmed by qPCR and Western blot in vitro and in vivo. PY34 and SCH900776 induced the downregulation of c-Myc mRNA and protein expression in a time-dependent manner in both MV411 and Z138 cells (Fig. 3e, f, h, i and Supplementary Fig. S4a, b). The downregulation of c-Myc protein and mRNA levels was significantly positively correlated (Fig. 3g, j), implying that transcriptional changes are the main reason behind the reduced protein levels. Furthermore, a similar effect was observed in PY34-treated MV411 xenograft tumors (Fig. 3k–m). Treatment with the proteasome inhibitor PS-341 did not reverse the c-Myc protein downregulation (Supplementary Fig. S5a), and PY34 treatment did not change the half-life of c-Myc upon CHX treatment (Supplementary Fig. S5b). Nevertheless, serine 62 phosphorylation (pS62) (which stabilizes c-Myc) was increased, and threonine 58 phosphorylation (pT58) (which promotes c-Myc degradation) was decreased in a time-dependent manner upon PY34 treatment (Supplementary Fig. S5c–e). We concluded that the PY34-induced decrease in c-Myc expression was independent of protein degradation but was achieved mainly through transcriptional regulation.
a–d RNA-seq data of Z138 and MV411 cells with or without PY34 treatment were analyzed. The commonly altered genes were selected (a), and the enriched signaling pathways were analyzed (b). Gene set enrichment analysis further confirmed that the c-Myc-changed gene signatures (c) and gene list (d) were downregulated significantly. e–m c-Myc downregulation was confirmed by qPCR and Western blot in MV411 (e, f) and Z138 cells (h, i) and in MV411 xenografts (k, l). The correlation between the downregulation of c-Myc protein and mRNA levels in MV411 (g), Z138 (j), and MV411 xenografts (m) was analyzed. *P < 0.05; **P < 0.01, versus vehicle group, unpaired two-tailed Student’s t test.
c-Myc plays an important role in PY34-induced anticancer effects
The above data showed that c-Myc downregulation is a common change in the CHK1i-sensitive cells tested. MV411 cells stably overexpressing c-Myc were established (Fig. 4a). Ectopic expression of c-Myc partially rescued the G1 phase arrest and growth delay caused by PY34 (Fig. 4b–d). PY34 could also induce c-Myc downregulation in BaF3 cells expressing either mutant FLT3-ITD (Fig. 4e) or FLT3-D835V (Fig. 4f) constructs. The addition of IL-3 led to the nearly complete restoration of c-Myc expression in PY34-treated FLT3-ITD and FLT3-D835V cells. Accordingly, IL-3 significantly increased resistance to PY34 in cells expressing FLT3-ITD or FLT3-D835V (Fig. 4g). Similarly, PY34 could induce the downregulation of c-Myc in TF-1 cells expressing mutated FLT3-ITD, and GM-CSF could reverse c-Myc downregulation and cell growth inhibition (Supplementary Fig. S9). These data showed that c-Myc functions as a key downstream effector of PY34, and c-Myc downregulation may be correlated with CHK1i efficacy.
Ectopic overexpression of c-Myc in MV411 cells (a) partially reversed G1 arrest (b, d) and cell viability (c). PY34 induced the change in c-Myc expression in wild-type BaF3 cells and BaF3 with FLT3-ITD (e) and in BaF3 cells with FLT3-D835V (f) in the presence or absence of IL-3; changes in the IC50 values of PY34 in BaF3 cells with FLT3-ITD or FLT3-D835V (g) were also observed. *P < 0.05; **P < 0.01, versus vehicle group, unpaired two-tailed Student’s t-test.
c-Myc downregulation is correlated with the sensitivity of HM cell lines and primary cells to PY34 treatment
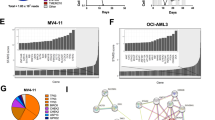
The levels of c-Myc, pS296-CHK1 and pS345-CHK1 as PD biomarkers of CHK1 inhibition were evaluated in four sensitive cell lines (Z138, MV411, CCRF-CEM, Ramos) and two insensitive cell lines (K562 and RPMI8226). The level of pS296-CHK1 decreased and that of pS345-CHK1 was increased in all these cell lines, indicating that CHK1 kinase activity was inhibited by PY34 in both sensitive and resistant cells. However, c-Myc protein levels were dramatically decreased in the sensitive cells but remained unchanged in the insensitive cells (Fig. 5a). Cell sensitivity to PY34 was closely correlated with the downregulation of c-Myc expression (Fig. 5b). The downregulation of c-Myc by CHK1i was mainly achieved at the transcriptional level (Fig. 3). The correlation between c-Myc mRNA downregulation by PY34 and cell sensitivity was further confirmed in 17 HM cell lines (Fig. 5c, d) and 39 primary HM cell samples (Fig. 5e–g). We found a significant negative correlation between the change in c-Myc expression and cell growth inhibition. These data suggested that c-Myc downregulation could be used as an indicator to predict the efficacy of CHK1i against HMs as early as possible.
Western blot (a) of CHK1, phosphorylated CHK1, and c-Myc in 6 HM cell lines treated with 200 nM PY34 for 8 h and the correlation (b) of c-Myc downregulation and the IC50 value of PY34. qPCR (c) of c-Myc downregulation in 17 HM cell lines treated with 200 nM PY34 for 8 h and the correlation of c-Myc downregulation and the IC50 value (d). IC50 values of PY34 in 39 HM patient samples (e), qPCR of c-Myc downregulation in 39 HM patient samples treated with 1000 nM PY34 for 8 h (f), and the correlation of c-Myc downregulation and the IC50 value (g). *P < 0.05; **P < 0.01, versus vehicle group, unpaired two-tailed Student’s t test.
Discussion
CHK1i has been used to treat blood cancers and solid cancers. Of all newly diagnosed cancers, 10.0% were estimated to be categorized as HMs. HMs are characterized by a deregulated DDR pathway, which makes HM cells more strongly dependent on CHK1 to allow adjustments in the cell cycle for DNA repair [22]. CHK1 is essential for normal B-cell development and the progression of lymphoma and leukemia [23, 24]. The high expression of CHK1 is positively correlated with shorter overall survival in AML patients with regard to cytarabine-based treatment [10]. CHK1i has been developed as a sensitizer for conventional therapeutics in patients with HMs [22, 25]. The GDSC database contains data from 161 HM cell lines and 718 solid tumor cell lines, where the CHK1is AZD7762 and SCH900776 showed obviously lower IC50 values in ALL, AML and multiple myeloma, the three major types of HMs (Fig. 1a, b, and Supplementary Fig. S1a, b). In addition, the DepMap database contains data on 58 HM and 505 solid tumor cancer cell lines, where HM cells are more dependent on CHK1 than solid tumor cells are (Fig. 1d). PY34 is a highly selective CHK1i in cells, as confirmed by the activity cluster of PY34 and six other CHK1is in clinical trials of HM lines (Supplementary Fig. S1d), which is in agreement with its high selectivity for kinase panel inhibition [18]. In 28 cancer cell lines, including 21 HM and 7 solid tumor cell lines, HM cells were more sensitive to PY34 than solid tumor cells (Fig. 1c). PY34 caused a dose-dependent induction of apoptosis in MV411, Z138, and CCRF-CEM cells (Fig. 2d, e, f) and exerted strong in vivo antitumor effects in a dose-dependent manner when used as a single agent in three xenograft models (Fig. 2g–o).
The mRNA expression of CHK1 in cells was extracted from CCLE, and the correlation with the IC50 value of the positive drugs AZD7762 and SCH90076 was analyzed (Supplementary Fig. S3a–f). There was no significant correlation between CHK1 expression and the IC50 value of the positive drugs AZD7762 and SCH90076 in total tumor samples, but there was only a weak correlation in HM samples. The mRNA expression of CHK1 in 12 HM cell lines was detected using qPCR (Supplementary Fig. S3g), and there was no significant correlation with the IC50 value of PY34 in these cell lines (Supplementary Fig. S3h).
PD biomarkers could provide information about the pharmacological effects of a drug on its target and even its desired biological effect. It is only slightly inconvenient for patients with HM to provide sufficient samples for PD biomarker research. Transcriptome-wide gene expression profiling has proven useful to improve our understanding of the underlying molecular mechanisms of drugs and drug sensitivity. Although PY34 arrested MV411 cells in G1 phase and Z138 cells in S phase (Fig. 2a, b), the RNA-seq data of MV411 and Z138 cells were analyzed, and the results showed that the expression of 208 genes showed similar changes in both cell lines upon treatment with PY34. The expression of genes targeted by the oncogenic transcription factor c-Myc were significantly downregulated (Fig. 3a–d), and these results were further confirmed by qPCR and Western blot in vitro and in vivo (Fig. 3e-m). The proteasome inhibitor PS-341 could not rescue the downregulated expression of c-Myc protein (Supplementary Fig. S5a). The half-life of the c-Myc protein was even slightly extended after PY34 treatment in the CHX-treated cells (Supplementary Fig. S5b), which may be caused by the increase in pS62 (which stabilizes c-Myc) and the decrease in pT58 (which promotes c-Myc degradation) (Supplementary Fig. S5c–e). Proteomic analysis revealed (i) higher protein levels of c-Myc in SCLC cells sensitive to the CHK1i LY2606368 and (ii) decreased expression of c-Myc after LY2606368 treatment in sensitive cells, but the underlying mechanisms were not studied [26]. Similarly, c-Myc protein levels in the human B-cell lymphoma lines Akata and BJAB, which are sensitive to CHK1is, declined after LY2606368 treatment, but no change was observed in the relatively resistant DG75, Kem1, and Raji cell lines [27]. A highly significant correlation between c-Myc and CHK1 mRNA levels in Burkitt lymphoma cells has been reported [27]. However, we did not find a significant correlation between c-Myc and CHK1 mRNA levels in the CCLE database, which contains data from 1019 cancer cell lines (Supplementary Fig. S6). It was reported that p53 represses c-Myc transcription [28]. Here, p53 knockdown had a slight effect on c-Myc expression and PY34-induced c-Myc downregulation (Supplementary Fig. S7), implying that the mechanism underlying the PY34-induced decrease in c-Myc transcription in HM cells is dependent on cell type.
In addition, the effect of CHK1 knockdown on c-Myc expression was investigated. The results showed that CHK1 expression was knocked down by ~50%, and c-Myc expression changed significantly in Z138 and MV411 cells but not in K562 cells (Supplementary Fig. S8). However, the effect of CHK1 knockdown on c-Myc downregulation was not as pronounced as that of CHK1i treatment. There may be two reasons: (i) the efficacy of CHK1 knockdown is limited to ~50%; and (ii) CHK1i may have other off-target inhibitors that synergize with CHK1 inhibition. The approval of the first kinase inhibitor, Gleevec, in 2001 ushered in a paradigm shift for oncological treatment. To date, over 60 kinase inhibitors have been approved by the FDA. However, there are no drugs that target a single kinase; rather, the available drugs largely inhibit multiple kinases.
It is known that c-Myc indirectly induces CHK1 mRNA and protein expression [27, 29]. Here, we also found that c-Myc overexpression could (i) increase CHK1 protein levels and partially reverse PY34-induced CHK1 downregulation (Fig. 4a) and (ii) partially reverse PY34-induced inhibition and G1 arrest (Fig. 4b, c). Interestingly, the DepMap database showed that HM cells were more dependent on c-Myc than solid tumor cells were (Supplementary Fig. S10a), similar to CHK1 cells (Fig. 1c, d). There were codependent relationships between c-Myc and CHK1 in HM cells but not in solid tumor cells (Supplementary Fig. S10b–d).
The reported results for pS345-CHK1 induction and pS296-CHK1 inhibition as PD biomarkers of CHK1i activity were reproduced in all PY34-treated HM cells (Fig. 5a), indicating that the change in CHK1 phosphorylation could be a mechanism of action biomarker for PY34. However, it could not be a PD biomarker related to CHK1i efficacy because there was no difference in PY34-treated sensitive and resistant cells. Importantly, PY34-induced downregulation of c-Myc protein levels was significantly negatively correlated with the IC50 value in six hematologic tumor cell lines (Fig. 5b). We confirmed that CHK1is decrease c-Myc protein levels mainly via downregulation of transcription. The PY34-induced c-Myc mRNA downregulation was significantly negatively correlated with its IC50 value in 17 hematologic tumor cell lines (Fig. 5c, d). However, the basal level of c-Myc gene expression had no correlation with the IC50 values in these hematologic tumor cell lines (Supplementary Fig. S11a, b). There was no significant correlation between c-Myc expression extracted from CCLE and the IC50 value of the positive drugs AZD7762 and SCH90076 in HM samples, but there was only a weak correlation in solid or total tumor samples (Supplementary Fig. S11c–h). Then, the correlation between c-Myc downregulation and drug sensitivity was tested in HM PDC samples. Twenty of the 39 primary human leukemia cells were sensitive to PY34 with an IC50 value less than 1 μM, and the degree of c-Myc downregulation induced by 1 μM PY34 was significantly negatively correlated with the extent of proliferation inhibition (Fig. 5e–g).
Conclusion
In summary, we have identified that HMs are more sensitive to CHK1is than solid tumors are via bioinformatics analysis of the GDSC and DepMap databases integrated with our experimental confirmation of the CHK1i PY34. Furthermore, downregulation of c-Myc expression is significantly associated with drug sensitivity to CHK1is in HM cell lines and PDC samples. Our findings suggest that c-Myc downregulation could serve as a potential biomarker for the a priori identification of HMs that are likely to respond to monotherapy with CHK1i, which can be helpful for the selection of CHK1i-sensitive cancer patients; therefore, this phenomenon deserves further investigation in follow-up preclinical studies and human clinical trials.
References
Sanchez Y, Bachant J, Wang H, Hu F, Liu D, Tetzlaff M, et al. Control of the DNA damage checkpoint by chk1 and rad53 protein kinases through distinct mechanisms. Science. 1999;286:1166–71.
Sanchez Y, Wong C, Thoma RS, Richman R, Wu Z, Piwnica-Worms H, et al. Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science. 1997;277:1497–501.
Madoz-Gurpide J, Canamero M, Sanchez L, Solano J, Alfonso P, Casal JI. A proteomics analysis of cell signaling alterations in colorectal cancer. Mol Cell Proteom. 2007;6:2150–64.
Sarmento LM, Povoa V, Nascimento R, Real G, Antunes I, Martins LR, et al. CHK1 overexpression in T-cell acute lymphoblastic leukemia is essential for proliferation and survival by preventing excessive replication stress. Oncogene. 2015;34:2978–90.
Boudny M, Zemanova J, Khirsariya P, Borsky M, Verner J, Cerna J, et al. Novel CHK1 inhibitor MU380 exhibits significant single-agent activity in TP53-mutated chronic lymphocytic leukemia cells. Haematologica. 2019;104:2443–55.
Cole KA, Huggins J, Laquaglia M, Hulderman CE, Russell MR, Bosse K, et al. RNAi screen of the protein kinome identifies checkpoint kinase 1 (CHK1) as a therapeutic target in neuroblastoma. Proc Natl Acad Sci USA. 2011;108:3336–41.
Tho LM, Libertini S, Rampling R, Sansom O, Gillespie DA. Chk1 is essential for chemical carcinogen-induced mouse skin tumorigenesis. Oncogene. 2012;31:1366–75.
Chen Z, Xiao Z, Chen J, Ng SC, Sowin T, Sham H, et al. Human chk1 expression is dispensable for somatic cell death and critical for sustaining G2 DNA damage checkpoint. Mol Cancer Ther. 2003;2:543.
Wang WT, Catto JWF, Meuth M. Differential response of normal and malignant urothelial cells to CHK1 and ATM inhibitors. Oncogene. 2014;34:2887–96.
David L, Fernandez-Vidal A, Bertoli S, Grgurevic S, Lepage B, Deshaies D, et al. CHK1 as a therapeutic target to bypass chemoresistance in AML. Sci Signal. 2016;9:ra90.
Meng Y, Chen CW, Yung MMH, Sun W, Sun J, Li Z, et al. DUOXA1-mediated ROS production promotes cisplatin resistance by activating ATR-Chk1 pathway in ovarian cancer. Cancer Lett. 2018;428:104–16.
Alsubhi N, Middleton F, Abdel-Fatah TM, Stephens P, Doherty R, Arora A, et al. Chk1 phosphorylated at serine345 is a predictor of early local recurrence and radio-resistance in breast cancer. Mol Oncol. 2016;10:213–23.
Fang Z, Gong C, Yu S, Zhou W, Hassan W, Li H, et al. NFYB-induced high expression of E2F1 contributes to oxaliplatin resistance in colorectal cancer via the enhancement of CHK1 signaling. Cancer Lett. 2018;415:58–72.
Mouw KW, Konstantinopoulos PA. From checkpoint to checkpoint: DNA damage ATR/Chk1 checkpoint signalling elicits PD-L1 immune checkpoint activation. Br J Cancer. 2018;118:933–5.
Lee J-M, Nair J, Zimmer A, Lipkowitz S, Annunziata CM, Merino MJ, et al. Prexasertib, a cell cycle checkpoint kinase 1 and 2 inhibitor, in BRCA wild-type recurrent high-grade serous ovarian cancer: a first-in-class proof-of-concept phase 2 study. Lancet Oncol. 2018;19:207–15.
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S, et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955–61.
Tsherniak A, Vazquez F, Montgomery PG, Weir BA, Kryukov G, Cowley GS, et al. Defining a cancer dependency map. Cell. 2017;170:564–76.
Tong L, Song P, Jiang K, Xu L, Jin T, Wang P, et al. Discovery of (R)-5-((5-(1-methyl-1H-pyrazol-4-yl)-4-(methylamino) pyrimidin-2-yl) amino)-3-(piperidin-3-yloxy) picolinonitrile, a novel CHK1 inhibitor for hematologic malignancies. Eur J Med Chem. 2019;173:44–62.
Chang K, Marran K, Valentine A, Hannon GJ. Creating an miR30-based shRNA vector. Cold Spring Harb Protoc. 2013;2013:631–5.
Shen A, Wang L, Huang M, Sun J, Chen Y, Shen YY, et al. c-Myc alterations confer therapeutic response and acquired resistance to c-Met inhibitors in MET-addicted cancers. Cancer Res. 2015;75:4548–59.
Liu H, Ai J, Shen A, Chen Y, Wang X, Peng X, et al. c-Myc alteration determines the therapeutic response to FGFR inhibitors. Clin Cancer Res. 2017;23:974–84.
Chamoun K, Borthakur G. Investigational CHK1 inhibitors in early stage clinical trials for acute myeloid leukemia. Expert Opin Investig Drugs. 2018;27:661–6.
Schuler F, Weiss JG, Lindner SE, Lohmuller M, Herzog S, Spiegl SF, et al. Checkpoint kinase 1 is essential for normal B cell development and lymphomagenesis. Nat Commun. 2017;8:1697.
Bryant C, Scriven K, Massey AJ. Inhibition of the checkpoint kinase Chk1 induces DNA damage and cell death in human leukemia and lymphoma cells. Mol Cancer. 2014;13:147.
Sampath D, Cortes J, Estrov Z, Du M, Shi Z, Andreeff M, et al. Pharmacodynamics of cytarabine alone and in combination with 7-hydroxystaurosporine (UCN-01) in AML blasts in vitro and during a clinical trial. Blood. 2006;107:2517–24.
Sen T, Tong P, Stewart CA, Cristea S, Valliani A, Shames DS, et al. CHK1 inhibition in small-cell lung cancer produces single-agent activity in biomarker-defined disease subsets and combination activity with cisplatin or olaparib. Cancer Res. 2017;77:3870–84.
Hoglund A, Nilsson LM, Muralidharan SV, Hasvold LA, Merta P, Rudelius M, et al. Therapeutic implications for the induced levels of Chk1 in Myc-expressing cancer cells. Clin Cancer Res. 2011;17:7067–79.
Ho JS, Ma W, Mao DY, Benchimol S. p53-dependent transcriptional repression of c-myc is required for G1 cell cycle arrest. Mol Cell Biol. 2005;25:7423–31.
Wang WJ, Wu SP, Liu JB, Shi YS, Huang X, Zhang QB, et al. MYC regulation of CHK1 and CHK2 promotes radioresistance in a stem cell-like population of nasopharyngeal carcinoma cells. Cancer Res. 2013;73:1219–31.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81673466, 81821005, 21772174, and 81172929), the “Personalized Medicines-Molecular Signature-based Drug Discovery and Development” and Strategic Priority Research Program of the Chinese Academy of Sciences (XDA12020214), the National Science and Technology Major Project of China (2018ZX09711002-011-015), and the Science and Technology Commission of Shanghai Municipality (16431902200).
Author information
Authors and Affiliations
Contributions
KLJ, YBZ, TL, JMY, and JL conceived and designed the study. KLJ, YBZ, LX, JL, KXZ, NS, JNL, JYL, MMZ, WBW, and YQX developed the methodology. KLJ, LXT, TW, HLW, XBH, GYX, TTJ, and WJK acquired the data (provided compounds and animals, enrolled and managed patients, provided facilities, etc.). KLJ, HLW, and YBZ analyzed and interpreted the data (e.g., statistical analysis, biostatistics, computational analysis). KLJ and YBZ wrote, reviewed, and/or revised the manuscript. XBH, GYX, TTJ, and WJK supplied administrative, technical, or material support (i.e., reporting or organizing data and constructing databases). AHG, YZH, YBZ, TL, JMY, and JL contributed to the study supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Jiang, Kl., Tong, Lx., Wang, T. et al. Downregulation of c-Myc expression confers sensitivity to CHK1 inhibitors in hematologic malignancies. Acta Pharmacol Sin 43, 220–228 (2022). https://doi.org/10.1038/s41401-021-00652-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-021-00652-1
Keywords
This article is cited by
-
Dual inhibition of CHK1/FLT3 enhances cytotoxicity and overcomes adaptive and acquired resistance in FLT3-ITD acute myeloid leukemia
Leukemia (2023)
-
Functions of lncRNA DUXAP8 in non-small cell lung cancer
Molecular Biology Reports (2022)