Abstract
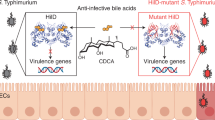
Microbiota generates millimolar concentrations of short-chain fatty acids (SCFAs) that can modulate host metabolism, immunity and susceptibility to infection. Butyrate in particular can function as a carbon source and anti-inflammatory metabolite, but the mechanism by which it inhibits pathogen virulence has been elusive. Using chemical proteomics, we found that several virulence factors encoded by Salmonella pathogenicity island-1 (SPI-1) are acylated by SCFAs. Notably, a transcriptional regulator of SPI-1, HilA, was acylated on several key lysine residues. Subsequent incorporation of stable butyryl-lysine analogs using CRISPR–Cas9 gene editing and unnatural amino acid mutagenesis revealed that site-specific modification of HilA impacts its genomic occupancy, expression of SPI-1 genes and attenuates Salmonella enterica serovar Typhimurium invasion of epithelial cells, as well as dissemination in vivo. Moreover, a multiple-site HilA lysine acylation mutant strain of S. Typhimurium was resistant to butyrate inhibition ex vivo and microbiota attenuation in vivo. Our results suggest that prominent microbiota-derived metabolites may directly acylate virulence factors to inhibit microbial pathogenesis in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data and materials that support the findings of this study are available from the corresponding author upon reasonable request.
References
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Cummings, J. H., Pomare, E. W., Branch, W. J., Naylor, C. P. & Macfarlane, G. T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28, 1221–1227 (1987).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Donohoe, D. R. et al. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 13, 517–526 (2011).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Voltolini, C. et al. A novel antiinflammatory role for the short-chain fatty acids in human labor. Endocrinology 153, 395–403 (2012).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Singh, N. et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 40, 128–139 (2014).
Macia, L. et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 6, 6734 (2015).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Chang, P. V., Hao, L., Offermanns, S. & Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl Acad. Sci. USA 111, 2247–2252 (2014).
Schulthess, J. et al. The short chain fatty acid butyrate imprints an antimicrobial program in macrophages. Immunity 50, 432–445.e7 (2019).
Byndloss, M. X. et al. Microbiota-activated PPAR-γ signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Rivera-Chávez, F. et al. Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host Microbe 19, 443–454 (2016).
Bronner, D. N. et al. Genetic ablation of butyrate utilization attenuates gastrointestinal Salmonella disease. Cell Host Microbe 23, 266–273.e4 (2018).
Gantois, I. et al. Butyrate specifically down-regulates Salmonella pathogenicity island 1 gene expression. Appl. Environ. Microbiol. 72, 946–949 (2006).
Lawhon, S. D., Maurer, R., Suyemoto, M. & Altier, C. Intestinal short-chain fatty acids alter Salmonella typhimurium invasion gene expression and virulence through BarA/SirA. Mol. Microbiol. 46, 1451–1464 (2002).
Groisman, Ea & Ochman, H. Cognate gene clusters govern invasion of host epithelial cells by Salmonella typhimurium and Shigella flexneri. EMBO J. 12, 3779–3787 (1993).
Hentchel, K. L. & Escalante-Semerena, J. C. Acylation of biomolecules in prokaryotes: a widespread strategy for the control of biological function and metabolic stress. Microbiol. Mol. Biol. Rev. 79, 321–346 (2015).
Yang, Y.-Y., Ascano, J. M. & Hang, H. C. Bioorthogonal chemical reporters for monitoring protein acetylation. J. Am. Chem. Soc. 132, 3640–3641 (2010).
Grammel, M. & Hang, H. C. Chemical reporters for biological discovery. Nat. Chem. Biol. 9, 475–484 (2013).
Hung, C.-C. C. et al. The intestinal fatty acid propionate inhibits Salmonella invasion through the post-translational control of HilD. Mol. Microbiol. 87, 1045–1060 (2013).
Sang, Y. et al. Acetylation regulating protein stability and DNA-binding ability of HilD, thus modulating Salmonella Typhimurium virulence. J. Infect. Dis. 216, 1018–1026 (2017).
Lostroh, C. P., Bajaj, V. & Lee, C. A. The cis requirements for transcriptional activation by HilA, a virulence determinant encoded on SPI-1. Mol. Microbiol. 37, 300–315 (2000).
Bajaj, V., Lucas, R. L., Hwang, C. & Lee, C. A. Co-ordinate regulation of Salmonella typhimurium invasion genes by environmental and regulatory factors is mediated by control of hilA expression. Mol. Microbiol. 22, 703–714 (1996).
Ellermeier, J. R. & Slauch, J. M. Adaptation to the host environment: regulation of the SPI1 type III secretion system in Salmonella enterica serovar Typhimurium. Curr. Opin. Microbiol. 10, 24–29 (2007).
Bajaj, V., Hwang, C. & Lee, C. A. hilA is a novel ompR/toxR family member that activates the expression of Salmonella typhimurium invasion genes. Mol. Microbiol. 18, 715–727 (1995).
Lostroh, C. P. & Lee, C. A. The HilA box and sequences outside it determine the magnitude of HilA-dependent activation of PprgH from Salmonella pathogenicity island 1. J. Bacteriol. 183, 4876–4885 (2001).
Eichelberg, K. & Galan, J. E. Differential regulation of Salmonella typhimurium type III secreted proteins by pathogenicity island 1 (SPI-1)-encoded transcriptional activators InvF and HilA. Infect. Immun. 67, 4099–4105 (1999).
Boddicker, J. D., Knosp, B. M. & Jones, B. D. Transcription of the Salmonella invasion gene activator, hilA, requires HilD activation in the absence of negative regulators. J. Bacteriol. 185, 525–533 (2003).
Christensen, D. G. et al. Identification of novel protein lysine acetyltransferases in Escherichia coli. MBio 9, e01905–e01918 (2018).
Sturm, A. Bistable Regulation of ttss-1 Genes in Salmonella Typhimurium. Doctoral Thesis, ETH Zürich (2011); https://doi.org/10.3929/ethz-a-006715280
Kim, D. E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Gattner, M. J., Vrabel, M. & Carell, T. Synthesis of ε-N-propionyl-, ε-N-butyryl-, and ε-N-crotonyl-lysine containing histone H3 using the pyrrolysine system. Chem. Commun. 49, 379–381 (2013).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Li, J. et al. Ligand-free palladium-mediated site-specific protein labeling inside gram-negative bacterial pathogens. J. Am. Chem. Soc. 135, 7330–7338 (2013).
Zhang, M. et al. A genetically incorporated crosslinker reveals chaperone cooperation in acid resistance. Nat. Chem. Biol. 7, 671–677 (2011).
Lin, S. et al. Site-specific incorporation of photo-cross-linker and bioorthogonal amino acids into enteric bacterial pathogens. J. Am. Chem. Soc. 133, 20581–20587 (2011).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, La RNA-guided editing of bacterial genomes using CRISPR–Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Drecktrah, D., Knodler, L. A., Ireland, R. & Steele-Mortimer, O. The mechanism of Salmonella entry determines the vacuolar environment and intracellular gene expression. Traffic 7, 39–51 (2006).
Barthel, M. et al. Pretreatment of mice with streptomycin provides a Salmonella enterica serovar Typhimurium colitis model that allows analysis of both pathogen and host. Infect. Immun. 71, 2839–2858 (2003).
Ellermeier, C. D., Ellermeier, J. R. & Slauch, J. M. HilD, HilC and RtsA constitute a feed forward loop that controls expression of the SPI1 type three secretion system regulator hilA in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 57, 691–705 (2005).
Hapfelmeier, S. et al. The Salmonella pathogenicity island (SPI)-2 and SPI-1 type III secretion systems allow Salmonella serovar Typhimurium to trigger colitis via MyD88-dependent and MyD88-independent mechanisms. J. Immunol. 174, 1675–1685 (2005).
Thijs, I. M. V. et al. Delineation of the Salmonella enterica serovar Typhimurium HilA regulon through genome-wide location and transcript analysis. J. Bacteriol. 189, 4587–4596 (2007).
Jacobson, A. et al. A gut commensal-produced metabolite mediates colonization resistance to Salmonella infection. Cell Host Microbe 24, 296–307.e7 (2018).
Sorbara, M. T. et al. Inhibiting antibiotic-resistant Enterobacteriaceae by microbiota-mediated intracellular acidification. J. Exp. Med. 216, 84–98 (2019).
Santiviago, C. A. et al. Analysis of pools of targeted Salmonella deletion mutants identifies novel genes affecting fitness during competitive infection in mice. PLoS Pathog. 5, e1000477 (2009).
Rangan, K. J., Yang, Y.-Y., Charron, G. & Hang, H. C. Rapid visualization and large-scale profiling of bacterial lipoproteins with chemical reporters. J. Am. Chem. Soc. 132, 10628–10629 (2010).
Peng, T. & Hang, H. C. Bifunctional fatty acid chemical reporter for analyzing S-palmitoylated membrane protein–protein interactions in mammalian cells. J. Am. Chem. Soc. 137, 556–559 (2015).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteom. 13, 2513–2526 (2014).
Court, D. L., Sawitzke, Ja & Thomason, L. C. Genetic engineering using homologous recombination. Annu. Rev. Genet. 36, 361–388 (2002).
Acknowledgements
We thank M. T. Mark, H. Zebroski and H. Molina at The Rockefeller Proteomics Resource Center for LC–MS analyses and butyrylated peptide synthesis. We thank Weill Cornell School of Medicine Electron Microscopy and Histology Core and Memorial Sloan-Kettering Cancer Center Pathology Core for assistance with histology sample preparation and scoring. We thank S. Lin and P. Chen (Peking University) for Salmonella amber codon expression plasmids, W. Jiang and L. Marraffini (Rockefeller University) for bacterial CRISPR–Cas9 plasmids, H. Andrews-Polymenis (Texas A&M College of Medicine) and M. McClelland (University of California, Irvine) for S. Typhimurium ΔhilA strain from NIAID, NIH: Salmonella enterica subsp. enterica, strain 14028S (serovar Typhimurium) single-gene deletion mutant library, J. Slauch (University of Illinois at Urbana-Champaign) for S. Typhimurium tetRA-hilD-3xFlag strain, C.-C. Hung and C. Altier (Cornell University) for providing S. Typhimurium strain 14028S and helpful discussions and J. Galan (Yale University) for helpful discussions. Z.J.Z. received support from the David Rockefeller Graduate Program. H.C.H. acknowledges support from NIH grants (R01GM087544 and R01AT007671) and the Lerner Trust.
Author information
Authors and Affiliations
Contributions
H.C.H. conceived the project, Z.J.Z. and V.A.P. performed experiments and data analysis, T.P. synthesized bmK, Z.J.Z. and H.C.H. wrote the paper, and all authors contributed to manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–5 and Figs. 1–14
Dataset 1
LFQ proteomic analysis of alk-3 labeling in S. Typhimurium. All significant hits are bold. Red, SPI-1 proteins. Blue, metabolic enzymes.
Rights and permissions
About this article
Cite this article
Zhang, Z.J., Pedicord, V.A., Peng, T. et al. Site-specific acylation of a bacterial virulence regulator attenuates infection. Nat Chem Biol 16, 95–103 (2020). https://doi.org/10.1038/s41589-019-0392-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0392-5
This article is cited by
-
Spatiotemporal and direct capturing global substrates of lysine-modifying enzymes in living cells
Nature Communications (2024)
-
Computational design and genetic incorporation of lipidation mimics in living cells
Nature Chemical Biology (2024)
-
Anti-infective bile acids bind and inactivate a Salmonella virulence regulator
Nature Chemical Biology (2023)
-
Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome
Nature Methods (2022)
-
Identification of an N-acetylneuraminic acid-presenting bacteria isolated from a human microbiome
Scientific Reports (2021)